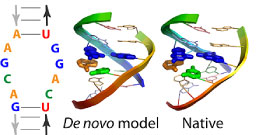
Our lab strives to model and design the RNA sequences that define and regulate living systems.
We develop algorithms to predict the structures and energetics of RNAs and RNA/protein interfaces with high accuracy. We test these ideas through community-wide blind trials; by enhancing NMR, crystallographic, and cryoelectron microscopy methods; and by designing new nanostructures.
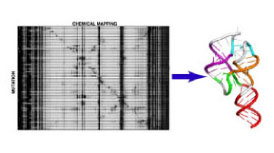
Complementary to our computational research, we are developing information-rich biochemical methods to model the myriad structures of non-coding RNAs that remain unknown. Current efforts focus on probing the extent of RNA structure and conformational change inside cells and viruses, in close collaboration with expert biologists at Stanford.